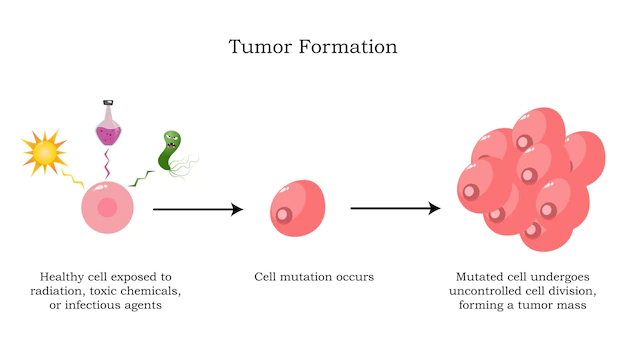
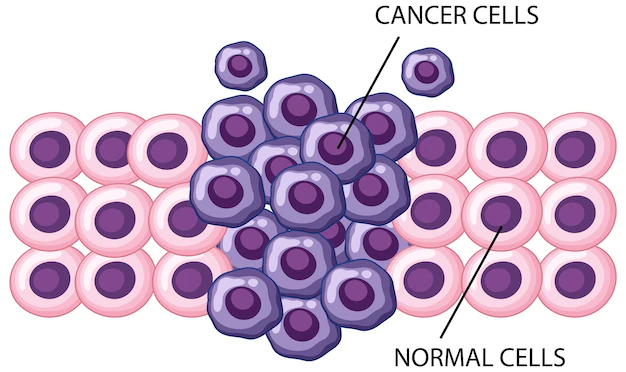
Mutagenic Impact Of Environmental Exposure – How It Induces Mutations In Human Cells
Mutagenic impact of environmental exposure damage DNA. Spontaneous chemical reactions such as hydrolysis may damage DNA, which causes nucleotide deamination.
Author:Suleman ShahReviewer:Han JuJul 19, 20221 Shares308 Views
As we age, somatic mutations develop in the DNA of our cells. As a consequence of copying DNA, these mutations are integrated during replication.
Mutagenic impact of environmental exposuredamage DNA. Spontaneous chemical reactions such as hydrolysis may damage DNA, which causes nucleotide deamination.
Balancing DNA damage induction and repair efficiency determines a cell's mutational landscape.
Mutagenic mechanisms cause most mutations that accumulate in healthy tissues throughout normal aging.
This theory might explain why aging is the leading risk factor for cancer. It is believed that smoking causes 80-90% of lung cancers, while UV radiation causes 86% of melanoma instances.
Environmental Genotoxicity Induced Mutations Detection
Many biological techniques were used to test the mutagenic potential of environmental exposure.
The use of reporter genes, such as LacZ in mouse models, has been used to research mutagenicity in a mammalian environment. Such an experimental technique is not conceivable in people. This gene encodes hypoxanthine-guanine phosphoribosyltransferase (HPRT), essential for synthesizing purine nucleotides.
Cells with HPRT activity also convert 6-thioguanine (6-TG) into a poisonous guanine analogue, which causes cell death. Sequencing cancer reporter genes like TP53 indicated the occurrence of CC > TT mutations in melanoma.
Sequencing Of Cancer Genomes
Mutation identification in complete genomes has become achievable with next-generation sequencing methods. Thousands of cancer exomes and genomes have been sequenced as part of critical consortium-based initiatives.
Traditional cancer genome sequencing approaches cannot identify mutagenic mechanisms active after the tumor starts and are stochastically present in a single or small fraction of tumor cells.
Mutational Signatures
Different mutagenic exposures may be disentangled over a cell's lifespan by finding recurring mutational patterns, or "mutational signatures," across cancer genomes. Individual mutation classes, such as single base substitutions, define these mutational signatures.
Different mechanisms may favor the straight 5' and 3' bases surrounding the altered base. Dimension reduction techniques may condense mutation spectra into a restricted collection of 96-trinucleotide signatures from mutation data.
In the recent decade, a rising number of mutational signatures have been established by studying increasing cancer genomes.
The biological cause of many signatures is unclear, while an underlying molecular link has been hypothesized for others. Among the molecular reasons are endogenous mechanisms that are active in all body cells and exposure to particular chemotherapeutic drugs.
Future signature extraction advancements will concentrate on the incorporation of additional genomic properties such as particular genomic regions and tissue-specific signatures.
Accumulation Of Mutations In Normal Cells
Normal cell genome sequencing may determine which mutagenesis mechanisms currently function in non-malignant cells. Because of the stochastic nature of mutation accumulation and the polyclonal architecture of most tissues, detecting somatic mutations in normal bulk tissue is challenging.
To get enough DNA from the parental cell, single stem/progenitor cells were expanded in vitro into clonal cultures. Whole genome amplification (WGA) based approaches are less appropriate for detecting mutations in samples with a low mutation load, such as normal cells.
New potential approaches for studying somatic mutations in single cells that address amplification-induced artifacts have been developed.
Primary template-directed amplification (PTA) is based on using exonuclease-resistant amplification terminators, while duplex-sequencing allows for the sequencing of two complementary DNA strands.
Errors during sequencing may be addressed by comparing the sequences since the two strands do not share them.
Normal Cell DNA Mutagenesis
During life, the genomes of healthy cells continually acquire mutations. Some mechanisms operate in most tissues in a clock-like fashion, generating mutation accumulation at a consistent pace that is constant within a tissue.
The mutation spectra and signatures show more dramatic variance across organs, implying tissue-specific activity of mutagenic mechanisms.
Half of all people investigated in colonic crypts had a particular mutational signature, defined by T > N single base changes in an ANNT context.
On rare occasions, E. coli has been found in the genomes of bladder, neuroendocrine, and head and neck malignancies. This mutational signature in these malignancies is most likely due to colibactin exposure.
UV-induced mutations have also been found in skin-resident lymphocytes and, on rare occasions, T-cell lymphoma.
In Vitro Assays Mutagenicity
Cell culture tests may also be used to investigate the mutational characteristics of environmental components. Cells undergo background mutagenesis, which seems to be driven by oxidative stress, and mutation accumulation due to genotoxic exposure during culture.
Groundbreaking research investigated the mutational effect of 79 distinct environmental genotoxins, offering a resource for causally linking mutational fingerprints to particular mutagenic exposures.
Treatment-Induced Signatures
Treatment might potentially harm noncancerous cells. This may lead to the buildup of DNA mutations in normal tissues, which might have long-term consequences.
Loss of a particular DNA repair activity may change the mutational profile generated by an environmental mutagen. Temozolomide exposure, for example, has been linked to two distinct mutational signatures.
SBS17a/b mutations may be caused by the chemotherapy medication 5-FU/capecitabine, which causes T > G alterations in a CpTpT context.
This hallmark has also been found in the genomes of untreated esophageal and stomach cancers. While the precise process is uncertain, it has been suggested that it is produced by including oxidized guanine residues opposite adenine in the DNA during replication.
MUTAGENIC AGENTS
Environmentally Induced Mutation Topographies
In vitro investigations may establish a link between a specific environmental mutagen and a mutational signature seen in the genomes of healthy and cancerous cells.
Mutations include extra information that may aid in determining the underlying process, such as genomic distribution and strand asymmetry. These traits combine to generate a "mutational topography," which may provide more insight into the origin of mutational signatures.
One of the major conformation-changing processes that occur in DNA is its transcription into RNA. When RNA polymerase II comes across a blocking DNA lesion, it is unable to progress and will stall.
RNA Pol II recruits transcription-coupled nucleotide excision repair to continue transcription-coupled nucleotide excision repair (TC-NER). Only in actively transcribed areas of the genome can TC-NER occur.
This preferential repair of the transcribed strand causes mutational strand asymmetry. This asymmetry bias may reveal which nucleotide of the mutant base pair was first damaged.
Following brief mutagenic exposures, DNA replication may result in an asymmetric distribution of mutations across the Watson and Crick strands.
The mutagenicity of such lesions may be affected by the activity of various polymerases on the leading and lagging strands. The segregation of strand lesions was initially discovered in mice given the highly mutagenic chemical diethylnitrosamine (DEN), which causes liver cancer.
Beyond the fundamental 5' and 3' flanking bases, cancer mutagenesis mechanisms may show a distinct preference for a specific setting. Assessing an additional base on each side of the mutation results in 1,536 different pentanucleotide categories.
These mutations are sufficiently particular that they may be used to predict tumor tissue type. Because nucleotide excision repair cannot reach transcription factor-binding sites in melanomas, they are enriched for mutations.
Environmental Genotoxins As A Cancer Driver
Environmental genotoxins can cause many somatic mutations, often identifiable by the mutational fingerprints they leave in cell genomes. Such a mutational signature does not always imply that the exposure had a role in carcinogenesis.
Because carcinogenesis requires mutations in specific driver genes, environmental mutagens' evidence of driver gene activation might be utilized to gather more proof that 5.3% of the colorectal cancer-causing APC gene mutations include the extended motif indicative of colibactin-induced alterations.
Despite the well-established significance of driver mutations in cancer, current sequencing studies have shown that many cancer driver mutations may be found in normal tissues without becoming malignant.
Oncogenic BRAF V600E mutations have been discovered in 83% of cutaneous naevi (common moles). Mutations in traditional cancer driver genes, such as NOTCH1 and TP53, may be found in 18-32% of tiny clonal populations of skin cells.
These mutations may be responsible for the clonal growth of these driver-containing cells, although the tissue is otherwise normal. Deep sequencing of esophageal epithelium indicated the occurrence of clonal expansions similarly.
People Also Ask
What Are The Harmful Effects Of Mutagens?
Mutagenic substances, which may jeopardize the integrity of the genetic code by generating DNA mutations, are a severe hazard to human health.
They have long been linked to various genetically inherited disorders, including cancer, aging, and neurodegenerative diseases such as Alzheimer's.
What Is Mutagen Exposure?
Examples of mutagens are tobacco products, radioactive compounds, x-rays, ultraviolet radiation, and a broad range of chemicals. Mutagen exposure may result in DNA alterations that cause or contribute to certain diseases.
What Are Some Environmental Mutagens?
Radiations are examples of environmental mutagens. Mutagens are ionizing radiations such as X-rays, gamma rays, alpha particles, UV radiations, and radioactive decay.
How Do Environmental Mutagens Cause Mutations?
Environmental mutagens are chemical and physical factors found in the environment that cause genetic mutations or raise mutation rates in humans. Most mutagens are carcinogenic in humans or have genotoxic effects on the following generation through germ cells.
Conclusion
Cancer is uncommon in humans at reproductive age, owing to evolutionary pressure. If the tissue is still functional, there is no need to prevent clonal expansions carrying driver mutations.
However, aging and environmental mutagens may increase mutation levels and change clonal dynamics. These extra activities may be sufficient to amass the additional traits and reach the "tipping point" when cells become cancerous.
The time has come to analyze which environmental genotoxins might cause cancer and how thoroughly.

Suleman Shah
Author
Suleman Shah is a researcher and freelance writer. As a researcher, he has worked with MNS University of Agriculture, Multan (Pakistan) and Texas A & M University (USA). He regularly writes science articles and blogs for science news website immersse.com and open access publishers OA Publishing London and Scientific Times. He loves to keep himself updated on scientific developments and convert these developments into everyday language to update the readers about the developments in the scientific era. His primary research focus is Plant sciences, and he contributed to this field by publishing his research in scientific journals and presenting his work at many Conferences.
Shah graduated from the University of Agriculture Faisalabad (Pakistan) and started his professional carrier with Jaffer Agro Services and later with the Agriculture Department of the Government of Pakistan. His research interest compelled and attracted him to proceed with his carrier in Plant sciences research. So, he started his Ph.D. in Soil Science at MNS University of Agriculture Multan (Pakistan). Later, he started working as a visiting scholar with Texas A&M University (USA).
Shah’s experience with big Open Excess publishers like Springers, Frontiers, MDPI, etc., testified to his belief in Open Access as a barrier-removing mechanism between researchers and the readers of their research. Shah believes that Open Access is revolutionizing the publication process and benefitting research in all fields.

Han Ju
Reviewer
Hello! I'm Han Ju, the heart behind World Wide Journals. My life is a unique tapestry woven from the threads of news, spirituality, and science, enriched by melodies from my guitar. Raised amidst tales of the ancient and the arcane, I developed a keen eye for the stories that truly matter. Through my work, I seek to bridge the seen with the unseen, marrying the rigor of science with the depth of spirituality.
Each article at World Wide Journals is a piece of this ongoing quest, blending analysis with personal reflection. Whether exploring quantum frontiers or strumming chords under the stars, my aim is to inspire and provoke thought, inviting you into a world where every discovery is a note in the grand symphony of existence.
Welcome aboard this journey of insight and exploration, where curiosity leads and music guides.
Latest Articles
Popular Articles